It maybe a less known fact that the lower cutoff of PCR cleanup and other DNA minipreps may be dialed in by appropriately diluting the binding buffer with water. PCR cleanup kits usually don’t bind fragments below 50-100 bp, depending on the manufacturer. Using water, dilution of the chaotropic salts in the binding buffer sets this limit higher. DNA smaller than the new limit runs through the coloumn, while higher MW DNA adsorbs as before. This can sometimes save the effort of gel purification.
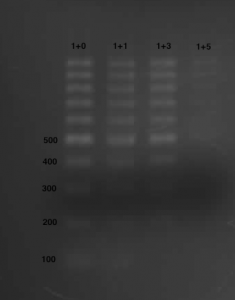
For the Macherey-Nagel Nucleospin this is described in the manual (pp 8-9). I found cutoff shifting works reasonably well also with the Qiaquick PCR cleanup kit of Qiagen. As illustrated with a 100bp DNA ladder below, diluting 1 parts binding buffer with 1 part water doesn’t change much, but 1+3 removes the 100 bp band without any harm to the higher MW bands. 1+5 removes bands below 500 bp, but here the loss of material seems prohibitive.
![David Molnar [Update:, PhD]](https://www.srcf.ucam.org/~dm516/wp-content/themes/twentyeleven/images/headers/chessboard.jpg)